Only accessible information is useful: insights from gradient-mediated patterning
Mikhail Tikhonov, Shawn C. Little, and Thomas Gregor. Roy. Soc. Open Sci. 2: 150486 (2015).
Abstract
Information theory is gaining popularity as a tool to characterize performance of biological systems. However, information is commonly quantified without reference to whether or how a system could extract and use it; as a result, information-theoretic quantities are easily misinterpreted. Here, we take the example of pattern-forming developmental systems which are commonly structured as cascades of sequential gene expression steps. Such a multi-tiered structure appears to constitute sub-optimal use of the positional information provided by the input morphogen because noise is added at each tier. However, one must distinguish between the total information in a morphogen and information that can be usefully extracted and interpreted by downstream elements. We demonstrate that quantifying the information that is accessible to the system naturally explains the prevalence of multi-tiered network architectures as a consequence of the noise inherent to the control of gene expression. We support our argument with empirical observations from patterning along the major body axis of the fruit fly embryo. We use this example to highlight the limitations of the standard information-theoretic characterization of biological signalling, which are frequently de-emphasized, and illustrate how they can be resolved.
PDF. Arxiv version.
Supplement.
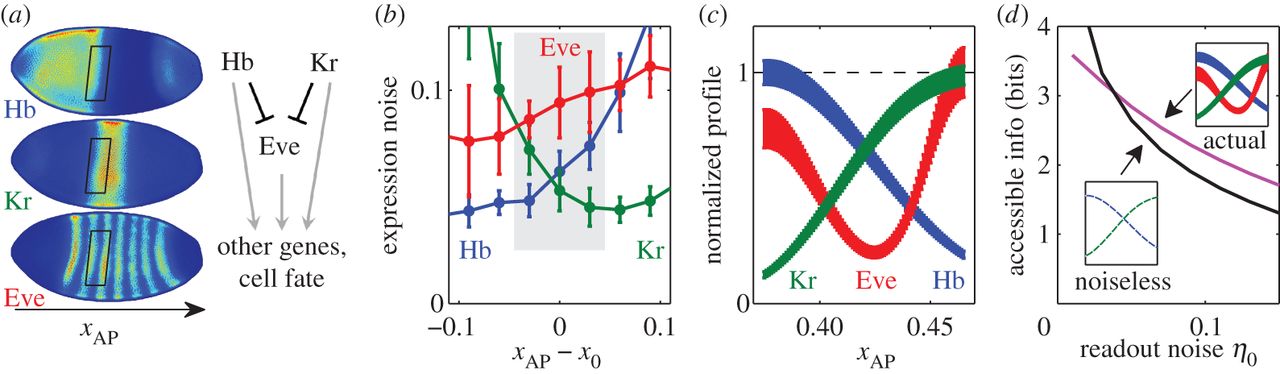
Figure 4 from the paper: (a) Immunostaining of three AP axis patterning genes in the same embryo. Rather than specifying cell fate directly, the ‘gap genes’ such as hunchback (Hb; top) and Krüppel (Kr; middle) control ‘pair-rule’ genes such as even-skipped (Eve, bottom). Both tiers regulate other genes further downstream. Boxes indicate the selected ROI, where at this time, Hb and Kr are the only relevant inputs to Eve, as shown in the cartoon. (b) Within the ROI (shaded), Eve exhibits higher expression noise than either Hb or Kr. Expression noise computed as RMS difference between expression level of a nucleus and its immediate dorsal or ventral neighbour (see the electronic supplementary material), plotted against AP distance from the Hb/Kr boundary (denoted x0). Error bars are standard deviation over N=8 embryos. (c) Idealized morphogen profiles, restricted to the ROI. Profile shape obtained as smooth spline-fit to expression values and noise magnitudes calculated for the profiles of panel (a) after projection onto the AP axis. (d) For all but the lowest readout noise magnitude, joint accessible information content in the triplet (Hb, Kr, Eve) exceeds the accessible information provided by Hb and Kr alone, even in an extreme hypothetical case when they are rendered entirely noiseless.